Explore our in vitro
primary cell immunology
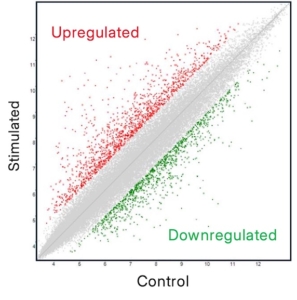
Gene expression: detecting activation before the protein appears
RT-qPCR targeting IFNG captures transcriptional activation; the cell’s commitment to producing IFN-γ before protein measurably accumulates in the supernatant. This makes it one of the most sensitive approaches for detecting early or low-magnitude T cell activation signals, and for profiling the transcriptional programmes driving the response. Well-validated primer sets and targeted pathway gene panels allow mechanistic resolution that protein-level assays cannot provide, including the co-regulation of downstream interferon-stimulated genes and upstream TCR signalling components.
Where RT-qPCR is targeted and hypothesis-driven, RNA-seq and other transcriptomic approaches offer more exploratory, offering whole-transcriptome resolution when the programme question extends beyond a defined target. The trade-offs are cost, analytical complexity, and turnaround time. For defined mechanistic questions, RT-qPCR is the more appropriate and cost-effective tool. For characterising an unfamiliar immune phenotype, better capturing biological variability, or mapping the transcriptional effects of a novel compound, -omics provides a breadth of resolution, and volume of data that targeted approaches simply can’t match.
It is important to note, however, that gene expression is an abstracted readout of immunological outcome. Upregulation of IFNG doesn’t truly confirm that protein was secreted, at what concentration, or that a functional immune response followed. mRNA abundance and protein output are related but not equivalent, and post-transcriptional regulation can decouple them. Gene expression data is therefore often most valuable as part of a broader analytical picture, providing mechanistic context and early signal detection rather than standing as sufficient evidence alone.
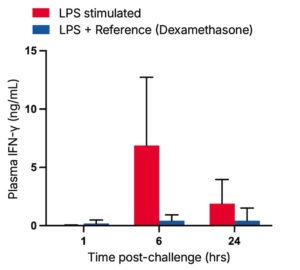
Soluble mediator quantification: the workhorse protein readout
ELISA remains the standard and one of our most common approaches for quantifying secreted IFN-γ in cell supernatants or biological matrices, as it’s sensitive, well-validated, and cost-effective at the scale required for multi-donor compound screens or dose-response studies. As an alternative, homogeneous immunoassay formats such as HTRF offer comparable sensitivity in miniaturised, wash-free formats that are also well suited to compound ranking and rapid screening workflows.
When the scientific question extends beyond IFN-γ alone, for example when looking at the broader cytokine landscape around T cell activation is beneficial, bead-based multiplex platforms allow simultaneous quantification of 20 or more analytes from a single sample volume. This is particularly valuable in early-stage and exploratory programmes, where profiling the full mediator context around activation can reveal pathway interactions, compensatory signalling, or off-target immune effects that a single-analyte approach wouldn’t detect.
A shared limitation across soluble mediator platforms is that they are population-level readouts. They quantify how much IFN-γ was released by a stimulated population. They do not reveal which cells were responsible for producing it.
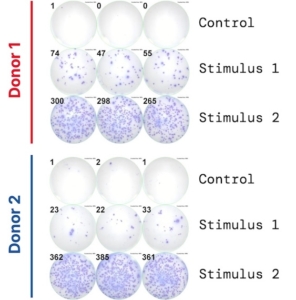
ELISpot and FluoroSpot: resolving how many cells respond
ELISpot addresses that limitation at the single-cell level, by counting individual IFN-γ secreting cells, as opposed to measuring supernatant concentrations. Each spot on the capture membrane represents a single cell that secreted IFN-γ in response to the applied stimulus. From a defined number of input PBMCs, the proportion of cells in a given donor’s immune repertoire capable of secreting IFN-γ under the experimental conditions is derived. The detection sensitivity of this approach is exceptional, reliably resolving responses from as few as one responding cell in 100,000.
This frequency-based resolution is what establishes ELISpot as the platform of choice for immunogenicity assessment, vaccine response monitoring, and antigen-specific T cell biology; applications where the number of responding cells is as important as the magnitude of the cytokine signal.
ELISpot is, however, a specialist platform with a narrow analyte scope of often one, or at most a small number of mediators per assay. FluoroSpot and multi-colour assays somewhat expand the number of analytes that can be detected simultaneously from the same well. This allows, for example, IFN-γ and IL-4 to be distinguished in the same population, or cytotoxic and regulatory responses to be quantified in parallel. In the right experimental context this adds meaningful resolution, but does mean these platforms are best suited when the demonstrable secretion of a defined, small set of mediators answers the experimental question.
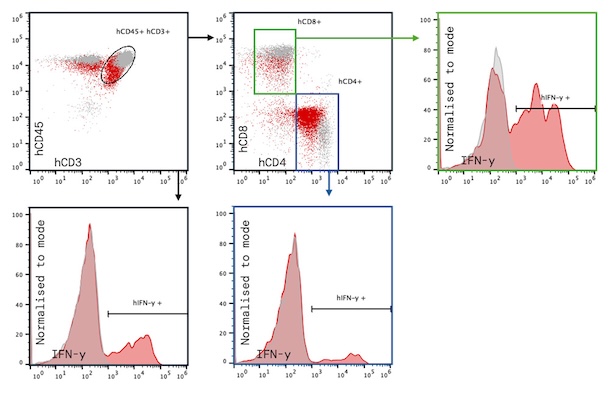
Flow cytometry and intracellular cytokine staining: adding immunophenotypic resolution
Flow cytometry with intracellular cytokine staining (ICS) answers a similar question to ELISpot with an added resolution layer; which specific cell subsets are producing IFN-γ, and the immunophenotypes of these cells.
By blocking intracellular secretion and staining for IFN-γ alongside a surface marker panel, ICS can identify the T cell populations responsible for cytokine production, CD4+ versus CD8+, effector versus memory, activated versus exhausted, and allows co-expression of multiple cytokines to be quantified in the same cell. A functional fingerprint of this kind, for example showing IFN-γ co-produced with TNF-α and IL-2, with a surface immunophenotype can provide deep mechanistic information and direct evidence of cell-specific effects or targeting.
The trade-off of this approach is that the sensitivity for low responding cell frequencies is generally lower than approaches like ELISpot. Panel designs also require validation and the approach requires greater time and cost investments than plate-based alternatives.
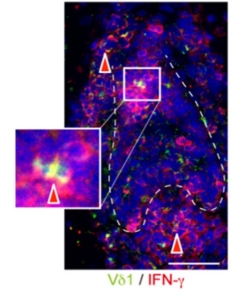
Representative immunofluorescence image sourced from Uchida et al. J Dermatol Sci 2020.
Spatial imaging: immune biology in solid tissue context
The platforms discussed so far resolve immune responses at the level of secreted mediators, cell frequencies, or cellular phenotype. What they don’t capture is the organisation of the immune response within solid tissue: the spatial relationships between immune populations and anatomical structures, or how immune activity maps onto tissue pathology.
Standard IHC and IF provide the entry point here, enabling single or dual marker detection on tissue sections interpreted alongside histopathological features. Structural changes such as alveolar thickening, crypt loss, or synovial hyperplasia contextualise immune cell density and distribution, and help link measurable responses to tissue level effects.
Multiplex immunostaining platforms such as Lunaphore COMET extend this to simultaneous multi-marker detection on the same section, resolving activated T cells, with additional immunophenotypes, and in spatial relationship to regulatory populations, stromal compartments, or target cells within intact tissue architecture.
These approaches do require solid tissue input, carry greater analytical complexity, and are lower throughput than plate-based methods (in bottom line terms, they can be expensive). For programmes where the architecture of the immune response within tissue is central to the research question, however, they provide a dimension of information that no other platform on this list can generate.
Platform selection as a scientific decision
It’s a common pitfall to reach for the most comprehensive endpoint by default. Measuring 20 cytokines simultaneously isn’t necessarily more informative than measuring one or two; in certain study designs it’s actually less so. Testing a large number of analytes without a clearly defined primary endpoint increases the multiple comparisons burden, requiring statistical corrections that reduce the confidence attributable to each individual readout. Across both solution-phase and tissue-based multiplex platforms, parallelised capture and sequential staining cycles introduce their own artefact risks, variable dynamic ranges, and technical complexity that single-plex or lower-order approaches avoid. The cost differential is also real and generally scales with analytical depth, so sophistication should be justified by the question, not applied by default.
The same principle extends to platform choice more broadly. Walking through five ways to measure T cell activation isn’t an argument for one approach over another, each is appropriate in a different context. In many programmes several will be used in sequence or for orthogonal validation, building resolution as the programme question demands it. It’s also worth noting that these platforms aren’t mutually exclusive. Samples can generally be aliquoted or fractionated across multiple readout approaches in the same experiment, allowing, for example, supernatant to go to ELISA while cells are taken forward to flow cytometry. This parallelisation is a genuine strength of a well-designed immunology study, though it carries its own opportunity costs in sample allocation, cost, and resolution that should be considered.
The right platform is ultimately the one that most efficiently answers the question the programme actually needs answered at that stage, within the constraints of available sample, budget, and timeline. At QIMA Life Sciences, this is treated as a scientific decision made in collaboration with client teams, working backwards from the programme question and the decision the data needs to enable.