Screening virtuel in silico SELNERGY
SELNERGY est un outil de screening in silico pour des applications biologiques. Il permet de modéliser de façon informatique les interactions entre des molécules chimiques (molécules actives ou inactives – ligands) et leurs cibles biologiques (protéines, enzymes, etc). SELNERGY est un outil de prospective puissant permettant de vous guider virtuellement dans le choix des pistes R&D à explorer.
Couplé à la ciblothèque GPDB et à une bibliothèque de 1 500 000 ligands actifs, le screening in silico SELNERGY permet d’identifier les molécules potentiellement actives pour une cible donnée ou inversement. SELNERGY donne une indication de leur(s) activité(s) biologique(s) et de leur(s) application(s) santé (pharmaceutique, cosmétique).
Cet outil de modélisation moléculaire se base à la fois sur une recherche de similarité de structure chimique 3D entre le composé étudié et les composés répertoriés dans les bases GPDB et de ligands actifs. SELNERGY permet de cribler plusieurs molécules ou cibles biologiques d’intérêt, de classer et de sélectionner les meilleurs candidats pour rationaliser les tests in vitro à réaliser ultérieurement.
Les deux approches de SELNERGY
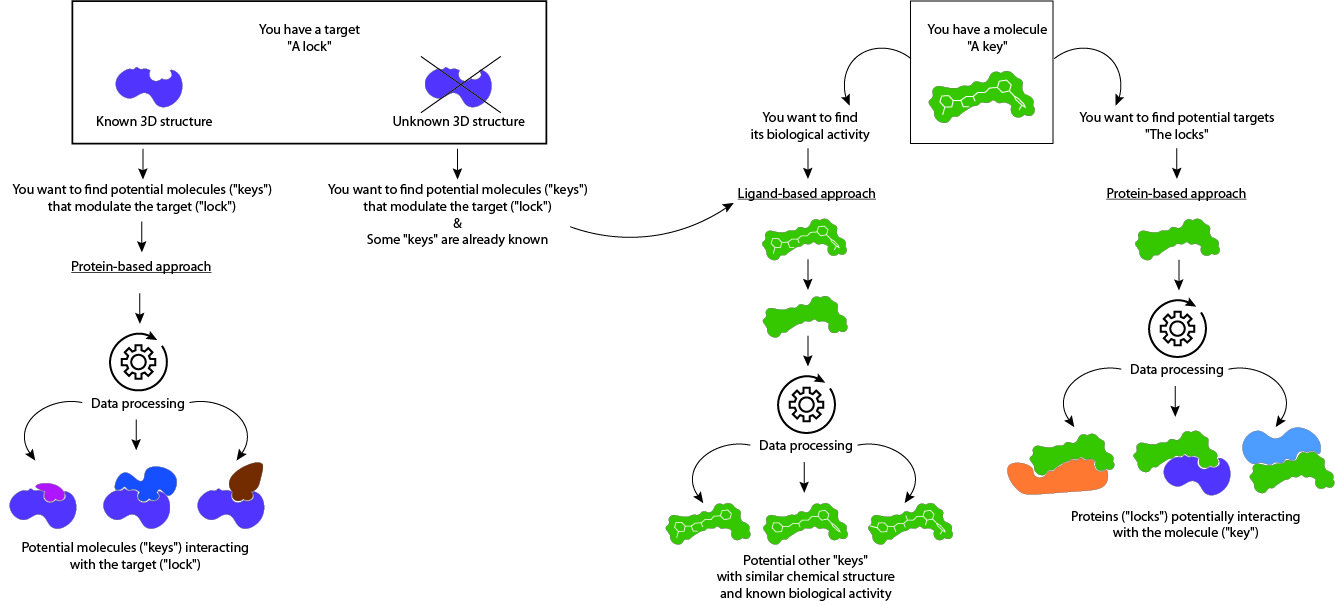
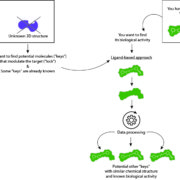
Ce screening in silico repose sur deux approches qui prennent en compte soit la structure de petites molécules actives (LBVS) soit la structure de la protéine cible (SBVS). Bien que l’une ou l’autre des approches peut-être choisie, les deux approches peuvent être utilisées de façon complémentaire :
-
Screening virtuel in silico basé sur le ligand (LBVS) : A partir d’une molécule de structure chimique connue, SELNERGY recherche dans les bases de données (GPDB et ligands actifs) des molécules aux structures 3D similaires, à la répartition de charges électrostatiques équivalente et à l’activité biologique connue. Les molécules obtenues par le logiciel sont classées par score de similarité. Les candidats ayant le meilleur score sont sélectionnés. Cette technique se base sur le postulat que si la structure chimique de deux molécules est similaire, leurs activités biologiques seront similaires. Aussi, il est possible de prédire l’activité biologique de la molécule d’intérêt et de préciser ses champs d’applications cosmétiques.
-
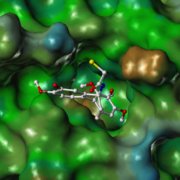
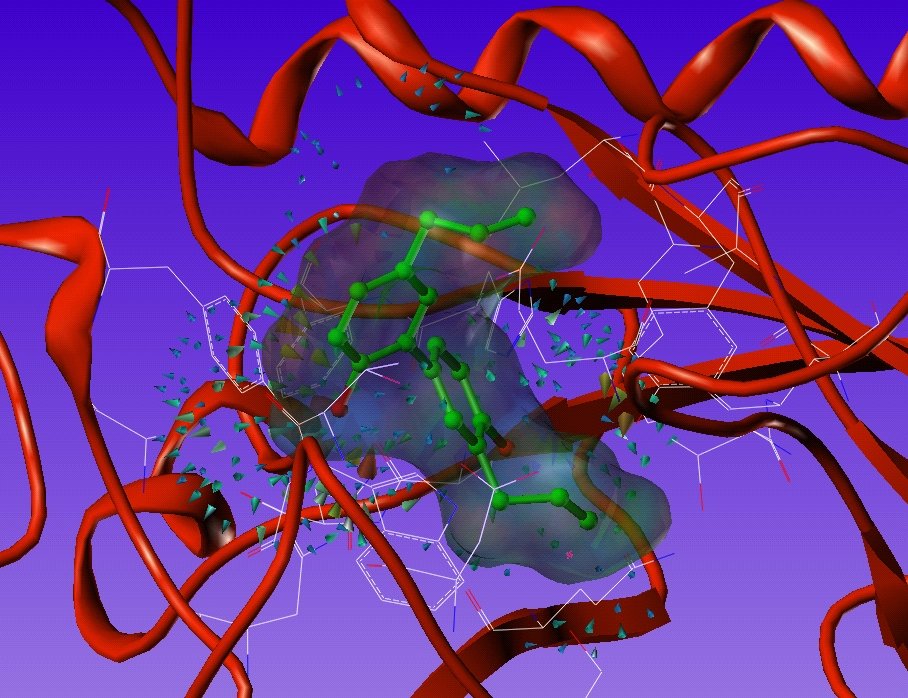
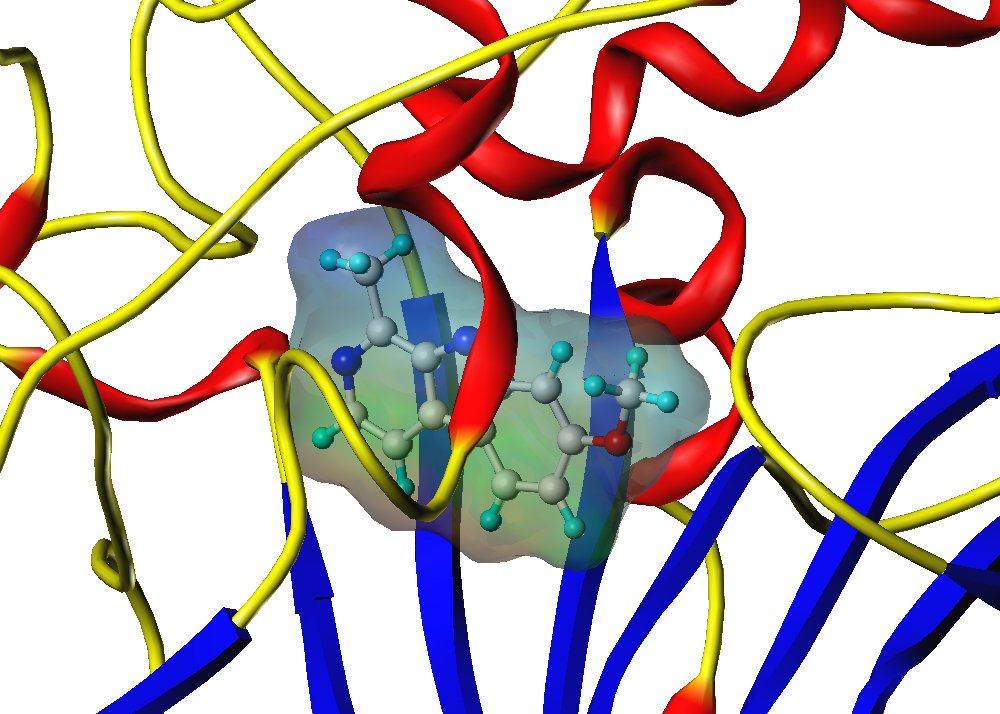
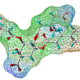
Screening virtuel in silico basé sur la structure de la protéine cible (SBVS) : A partir d’une cible biologique aux structure chimique et conformation 3D connues (protéine, enzyme), SELNERGY recherche dans les bases de données (GPDB et ligands actifs) des molécules capables de s’amarrer à la cible d’intérêt. Les molécules obtenues sont classées par score d’affinité et les meilleurs candidats sont sélectionnés. Cette méthode permet la découverte de composés potentiellement modulateurs de l’activité biologique de la cible.
De la même manière, à partir d’une molécule active à la structure chimique connue, SELNERGY recherche dans les bases de données des cibles biologiques potentielles. Cet « amarrage inverse » permet par conséquent de prédire l’activité biologique de la molécule d’intérêt et de là, ses champs d’application cosmétiques.